A medical image registration method based on network structure and circuit simulation
Authors: Li Chen, Yuxi Lian, Yi Guo*, Yuanyuan Wang, Thomas S. Hatsukami, Kristi Pimentel, Niranjan Balu, Chun Yuan
Abstract:
Image registration is an important research topic in the field of image processing. Applying registration to medical images allows multiple medical images to be strengthened and fused, which has practical value in disease detection, clinical assisted therapy, etc.
Image registration of tubular tree-like structures is a very difficult task. Common registration methods are usually unsuitable in vessel image registration. Area-based methods using pixel information for registration is sensitive to distortion and noise. Feature-based methods find correspondence between image features such as points, lines, and contours. However, vessel structures are lack of stable feature points, which may lead to false registration. Network-based methods are new approaches in vessel image registration. By transforming images to graph networks, nodes in the network are registered according to distance or feature. However, such methods are limited when facing topological missing and show high complexity in large size of images.
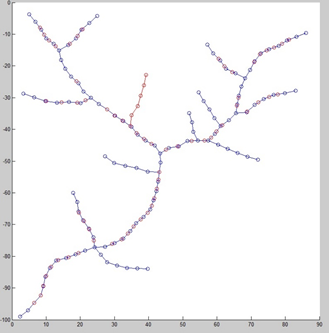
In this paper, we proposed a novel vessel image registration method based on network structure and circuit simulation. Firstly, vessel images were transformed to network graphs, with vessel radius as weight of each node. Then, based on regionally maximum Network Structure Index (NSI), a parameter to evaluate influence and connectivity of nodes, weighted network graphs were segmented to branches to reduce the calculation complexity. Enlightened by the similarity of graph network and circuit topology, a stable and efficient circuit simulation model was applied to register nodes in a branch. Finally, according to the circuit simulation model, properties of nodes were converted to a combination of resistance, inductance and voltage power source. Voltage of each node acquired from the power stimulation was used to describe features of nodes in branches, so that the comparison of voltage series of branches between registration image sets could be used to reflect node deviation of location in branches.
The proposed method was effective and efficient in vessel registration. The use of network structure tolerated noises and distortions, as vessel topology of image sets remained the same. In addition, NSI was also a regionally stable index determined by all its neighbors instead of a single node. A single erroneous node had little impact to the segmentation of the graph network, which was an efficient method in reducing calculation and finding topological missing. The circuit simulation was also a method stable against deviation. The voltages of the nodes were combined parameters from the weighted network and the voltage series was acquired from the average voltages during the power stimulation, effectively reflecting the feature of the network.
We tested our proposed method in artificial image sets and clinical image sets. The method was successful in finding the node deviation of location from the network in a reasonable time, showing the effective and efficient of our proposed method in registration.
Keywords:
image registration, network structure, circuit simulation, medical image
Abstract presented at bioimage informatics conference 2016
Full paper published in BMC Bioinformatics (SCIE, Impact Factor of 2.435)
Chen, Li, et al. "A vascular image registration method based on network structure and circuit simulation." BMC Bioinformatics 18.1 (2017): 229.
More information: http://clatfd.cn/a/165